Tutorial
Jelle, updated May 2020
updated Feb 2022 by Joran
This notebook shows how to do basic analysis with straxen, much like hax.minitrees.
For reference, here are some jargon terms which we will introduce below:
Context: Holds configuration on how to process
Dataframe or array: table of related information produced by a plugin.
Plugin: an algorithm that produces a dataframe
Data type: specification of which columns are in a dataframe.
Data kind: e.g. ‘events’ or ‘peaks’. Dataframes of the same kind have the same number of rows and can be merged.
[1]:
import numpy as np
import pandas as pd
# This just ensures some comments in dataframes below display nicely
pd.options.display.max_colwidth = 100
import straxen
[2]:
straxen.print_versions()
cutax is not installed
[2]:
| module | version | path | git | |
|---|---|---|---|---|
| 0 | python | 3.10.0 | /home/angevaare/miniconda3/envs/dev_strax/bin/python | None |
| 1 | strax | 1.1.7 | /home/angevaare/software/dev_strax/strax/strax | branch:master | db14f80 |
| 2 | straxen | 1.2.8 | /home/angevaare/software/dev_strax/straxen/straxen | branch:master | 024602e |
Setting up
First we load a strax context, much like hax.init(). A strax context contains all information on how to process: where to read what files from, what plugins provide what data, etc.
You can make a context yourselves using strax.Context, but straxen provides standardized contexts as well. Most future analyses will use such standardized contexts defined by analysis coordinators or straxen maintainers.
Unlike hax.init, you can have multiple active contexts, e.g. to load analysis and MC data, or compare data processed with different settings (we will see examples of this below).
[3]:
st = straxen.contexts.xenonnt_online()
Finding your data
Suposse we want to make a cS1/cS2 plot. We have to figure out which type of dataframes to load. A specific type of dataframe is also called a data type. (in hax these were called minitrees)
We can find this out automatically if we know (part of) the name of a field to load:
[4]:
st.search_field("cs1")
cs1 is part of corrected_areas (provided by CorrectedAreas)
cs1 is part of event_info (provided by EventInfo)
cs1 is used in CorrectedAreas.infer_dtype
cs1 is used in CorrectedAreas.compute
cs1 is used in EnergyEstimates.compute
cs1 is used in EnergyEstimates.cs1_to_e
It seems we’re after one of the data types called event_info or corrected_areas. In the current context, these are provided by plugins called EventInfo and CorrectedAreas, respectively (but this doesn’t concern us yet).
Additionally, we see the occurrences of cs1 of a field in EnergyEstimates and CorrectedAreas. This means that the field is used there directly
Let’s see what else is in these data types:
[5]:
st.data_info("event_info")
[5]:
| Field name | Data type | Comment | |
|---|---|---|---|
| 0 | time | int64 | Start time since unix epoch [ns] |
| 1 | endtime | int64 | Exclusive end time since unix epoch [ns] |
| 2 | cs1 | float32 | Corrected area of main S1 [PE] |
| 3 | cs2_wo_elifecorr | float32 | Corrected area of main S2 before elife correction (s2 xy correction + SEG/EE correction applied)... |
| 4 | cs2_wo_timecorr | float32 | Corrected area of main S2 before SEG/EE and elife corrections(s2 xy correction applied) [PE] |
| ... | ... | ... | ... |
| 106 | z_naive | float32 | Interaction z-position using mean drift velocity only (cm) |
| 107 | r_naive | float32 | Interaction r-position using observed S2 positions directly (cm) |
| 108 | r_field_distortion_correction | float32 | Correction added to r_naive for field distortion (cm) |
| 109 | z_field_distortion_correction | float32 | Correction added to z_naive for field distortion (cm) |
| 110 | theta | float32 | Interaction angular position (radians) |
111 rows × 3 columns
As you can see, event_info has a lot more information; let’s load that one. You can see from the documentation (TODO link) that event_info’s job is to merge the info from corrected_areas and other things.
Loading data
Next, you’ll want to select a run. The select_runs function will return a dataframe with all available runs; there is a separate tutorial on more advanced use of this. In this demo context, we only have high-level data for the run 180215_1029 available (and low-level data for another):
[ ]:
st.select_runs()
So lets’ take 021932.
To actually load data, you use get_df to get a pandas DataFrame, or get_array to get a numpy (structured) array. Let’s go with pandas for now:
[6]:
run_id = "021932"
# The seconds_range=[0,60] is an optional argument to prevent loading too much data at once
df = st.get_df(run_id, "event_info", seconds_range=[0, 60])
The first time you run this, it will take a moment: it has to actually process the data somewhat. We didn’t ship highest-level demo data with straxen: that would mean we’d have to constantly update the test data when the algorithms change.
You can also specify a list of runid’s instead of one run, and get the concatenated result back. Likewise, you can specify multiple data types (of the same kind) to load, and they will be merged for you.
Just like hax.minitrees.load, we got a dataframe back:
[7]:
st.show_config("event_info")
[7]:
| option | default | current | applies_to | help | |
|---|---|---|---|---|---|
| 0 | gain_model | <OMITTED> | (to_pe_model, ONLINE, True) | (afterpulses,) | PMT gain model. Specify as (model_type, model_config) |
| 1 | n_tpc_pmts | <OMITTED> | 494 | (afterpulses,) | Number of PMTs in TPC |
| 2 | LED_window_left | 50 | <OMITTED> | (afterpulses,) | Left boundary of sample range for LED pulse integration |
| 3 | LED_window_right | 100 | <OMITTED> | (afterpulses,) | Right boundary of sample range for LED pulse integration |
| 4 | baseline_samples | 40 | <OMITTED> | (afterpulses,) | Number of samples to use at start of WF to determine the baseline |
| ... | ... | ... | ... | ... | ... |
| 269 | recon_alg_included | (_mlp, _gcn, _cnn) | <OMITTED> | (s2_recon_pos_diff,) | The list of all reconstruction algorithm considered. |
| 270 | max_veto_window | 7200000000000 | <OMITTED> | (veto_intervals,) | Maximum separation between veto stop and start pulses [ns]. Set to be >> than the max duration o... |
| 271 | event_window_fields | (time, endtime) | <OMITTED> | (veto_proximity,) | Fields to determine where to look for overlaps for using this plugin in the events. The default ... |
| 272 | veto_proximity_window | 300000000000 | <OMITTED> | (veto_proximity,) | Maximum separation between veto stop and start pulses [ns] |
| 273 | time_no_aqmon_veto_found | 3600000000000 | <OMITTED> | (veto_proximity,) | If no next/previous veto is found, we will fill the fields time_to_previous_XX with this time. S... |
274 rows × 5 columns
[8]:
df.head()
[8]:
| time | endtime | cs1 | cs2_wo_elifecorr | cs2_wo_timecorr | cs2_area_fraction_top | cs2_bottom | cs2 | alt_cs1 | alt_cs2_wo_elifecorr | ... | alt_s2_y_mlp | x | y | z | r | z_naive | r_naive | r_field_distortion_correction | z_field_distortion_correction | theta | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1623144887003148092 | 1623144887005863030 | 2.659672 | 348.436493 | 370.887268 | 0.777278 | 82.604637 | 370.887268 | NaN | 34.098747 | ... | -23.990379 | -7.353356 | -15.314320 | -43.339104 | 16.988239 | -43.339104 | 16.190155 | 0.798083 | 0.007348 | -2.018448 |
| 1 | 1623144887006147772 | 1623144887008854250 | 13155.237305 | NaN | NaN | NaN | NaN | NaN | 4.279365 | NaN | ... | NaN | 45.517769 | 55.207436 | -0.189628 | 71.552284 | -0.189628 | 69.542831 | 2.009451 | 0.000000 | 0.881302 |
| 2 | 1623144887030319882 | 1623144887033327360 | 61723.960938 | 28599.917969 | 29029.152344 | 0.723225 | 8034.542969 | 29029.152344 | 19.304628 | 1338.243530 | ... | 53.898460 | -35.264309 | 55.141556 | -10.200156 | 65.453514 | -10.200156 | 61.628628 | 3.824886 | 0.744288 | 2.139771 |
| 3 | 1623144887054958322 | 1623144887057677700 | 76545.257812 | 1266.177612 | 1266.558594 | 0.159955 | 1063.966309 | 1266.558594 | 2.062342 | 17.207802 | ... | 2.884162 | 31.293451 | 9.913309 | -0.026945 | 32.826115 | -0.026945 | 37.803139 | -4.977024 | 0.000000 | 0.306784 |
| 4 | 1623144887127075472 | 1623144887129777490 | NaN | 5178.317383 | NaN | 0.749424 | NaN | NaN | NaN | NaN | ... | NaN | NaN | NaN | NaN | NaN | NaN | 61.323208 | NaN | NaN | 2.864871 |
5 rows × 111 columns
Let’s make a quick plot of the events we just loaded:
[9]:
df.plot.scatter("cs1", "cs2")
import matplotlib.pyplot as plt
plt.xscale("log")
plt.xlim(1, None)
plt.yscale("log")
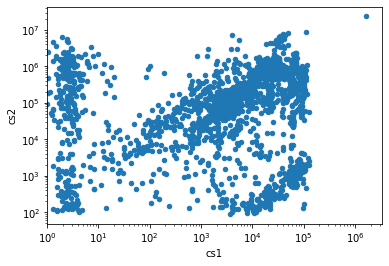
Since making a cS1, cS2 plot for a dataset is such a common task that straxen has a built-in method for it. There are other similar mini-analyses, such a waveform plotting, which we will see in action below.
[10]:
st.event_scatter(run_id, s=20, seconds_range=[0, 60])
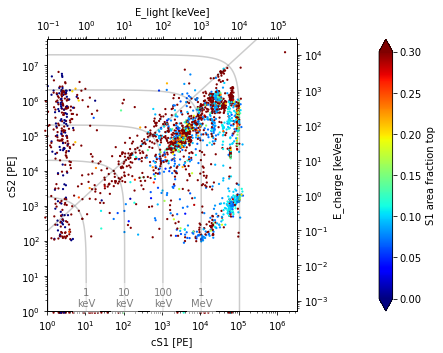
Can you guess what kind of data this is?
Waveform analysis
The peaks data type contains the sum waveform information:
[11]:
st.data_info("peaks")
[11]:
| Field name | Data type | Comment | |
|---|---|---|---|
| 0 | time | int64 | Start time since unix epoch [ns] |
| 1 | length | int32 | Length of the interval in samples |
| 2 | dt | int32 | Width of one sample [ns] |
| 3 | channel | int16 | Channel/PMT number |
| 4 | type | int8 | Classification of the peak(let) |
| 5 | area | float32 | Integral across channels [PE] |
| 6 | area_per_channel | ('<f4', (494,)) | Integral per channel [PE] |
| 7 | n_hits | int32 | Number of hits contributing at least one sample to the peak |
| 8 | data | ('<f4', (200,)) | Waveform data in PE/sample (not PE/ns!) |
| 9 | width | ('<f4', (11,)) | Peak widths in range of central area fraction [ns] |
| 10 | area_decile_from_midpoint | ('<f4', (11,)) | Peak widths: time between nth and 5th area decile [ns] |
| 11 | saturated_channel | ('i1', (494,)) | Does the channel reach ADC saturation? |
| 12 | n_saturated_channels | int16 | Total number of saturated channels |
| 13 | tight_coincidence | int16 | Channel within tight range of mean |
| 14 | max_gap | int32 | Largest gap between hits inside peak [ns] |
| 15 | max_goodness_of_split | float32 | Maximum interior goodness of split |
Notice the compound data types of the data, width and saturated_channel fields. Pandas does not support such types (well, it sort of does, but the resulting dataframes are quite inefficient), so we have to load this as a numpy array.
[12]:
peaks = st.get_array(run_id, "peaks", seconds_range=[0, 60])
type(peaks), peaks.dtype.names
[12]:
(numpy.ndarray,
('time',
'length',
'dt',
'channel',
'type',
'area',
'area_per_channel',
'n_hits',
'data',
'width',
'area_decile_from_midpoint',
'saturated_channel',
'n_saturated_channels',
'tight_coincidence',
'max_gap',
'max_goodness_of_split'))
Now we can plot peak waveforms:
[13]:
def plot_peak(p, t0=None, **kwargs):
n = p["length"]
if t0 is None:
t0 = p["time"]
plt.plot(
(p["time"] - t0) + np.arange(n) * p["dt"],
p["data"][:n] / p["dt"],
drawstyle="steps-mid",
**kwargs
)
plt.xlabel("Time (ns)")
plt.ylabel("Sum waveform (PE / ns)")
plot_peak(peaks[148])
plt.show()
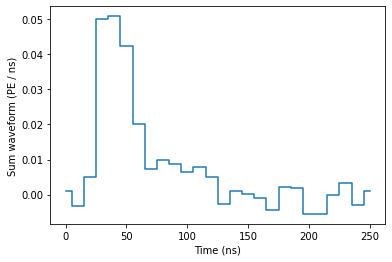
[14]:
def plot_peaks(main_i, n_before=0, n_after=0, label_threshold=0, legendloc="best"):
for i in main_i + np.arange(-n_before, n_after + 1):
p = peaks[mask][i]
label = None
if p["area"] > label_threshold:
label = "%d PE, %d ns dt" % (
p["area"],
p["dt"],
)
color = None
else:
color = "gray"
plot_peak(p, t0=peaks[main_i]["time"], label=label, color=color)
plt.ylim(0, None)
plt.legend(loc=legendloc)
plt.yscale("symlog")
# Find the largest peak close to 1e4 PE
mask = peaks["area"] < 1e4
i_of_largest_peak = np.argmax(peaks[mask]["area"])
plot_peaks(i_of_largest_peak, n_after=5, n_before=2, label_threshold=10, legendloc=(1.1, 0.0))
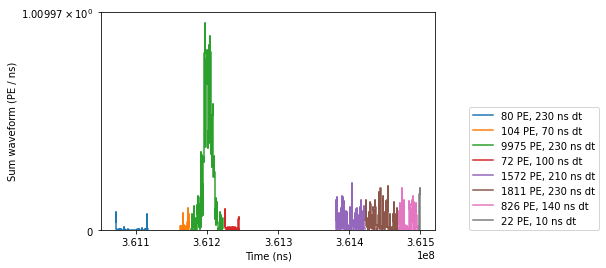
The abrupt termination of the S2 above is due to strax’s data reduction.
If you have access to the raw data (at least the records level) you can use straxen’s built-in waveform display. For example, try:
st.waveform_display(run_id, seconds_range=(0, 0.15))
(we didn’t evaluate this in the tutorial, as it creates a substantial amount of javascript, which would have made the notebook quite huge).
Configuration changes
As you can see in the above plot, we have many events high up in the TPC at low S1. Perhaps you want to get rid of them by increasing the ‘S1 coincidence requirement’, i.e. the number of PMTs that must see something before a peak is labeled as S1. Then, of course, you want to load the event-level data again to see if it worked.
First, we need to see which configuration option we have to change. Strax plugins declare what configuration they take and what other plugins they depend on, so this is not very difficult. We just ask which options with s1 in their name influence event_basics:
[29]:
st.show_config("event_basics", "s1*")[["option", "applies_to", "help", "current", "default"]]
[29]:
| option | applies_to | help | current | default | |
|---|---|---|---|---|---|
| 0 | s1_xyz_map | (corrected_areas,) | <OMITTED> | itp_map://resource://cmt://format://s1_xyz_map_{algo}?version=ONLINE&run_id=plugin.run_id&fmt=js... | |
| 1 | s1_risetime_area_parameters | (peaklet_classification,) | norm, const, tau in the empirical boundary in the risetime-area plot | <OMITTED> | (50, 80, 12) |
| 2 | s1_risetime_aft_parameters | (peaklet_classification,) | Slope and offset in exponential of emperical boundary in the rise time-AFT plot. Specified as (s... | <OMITTED> | (-1, 2.6) |
| 3 | s1_flatten_threshold_aft | (peaklet_classification,) | Threshold for AFT, above which we use a flatted boundary for rise timeSpecified values: (AFT bou... | <OMITTED> | (0.6, 100) |
| 4 | s1_max_rise_time_post100 | (peaklet_classification,) | Maximum S1 rise time for > 100 PE [ns] | <OMITTED> | 200 |
| 5 | s1_min_coincidence | (peaklet_classification,) | Minimum tight coincidence necessary to make an S1 | <OMITTED> | 2 |
| 6 | s1_optical_map | (event_pattern_fit,) | S1 (x, y, z) optical/pattern map. | <OMITTED> | XENONnT_s1_xyz_patterns_LCE_corrected_qes_MCva43fa9b_wires.pkl |
| 7 | s1_aft_map | (event_pattern_fit,) | Date drive S1 area fraction top map. | <OMITTED> | s1_aft_dd_xyz_XENONnT_Kr83m_41500eV_31Oct2021.json |
| 8 | s1_min_area_pattern_fit | (event_pattern_fit,) | Skip EventPatternFit reconstruction if S1 area (PE) is less than this | <OMITTED> | 2 |
| 9 | s1_risetime_area_parameters | (peaklet_classification_he,) | norm, const, tau in the empirical boundary in the risetime-area plot | <OMITTED> | (50, 80, 12) |
| 10 | s1_risetime_aft_parameters | (peaklet_classification_he,) | Slope and offset in exponential of emperical boundary in the rise time-AFT plot. Specified as (s... | <OMITTED> | (-1, 2.6) |
| 11 | s1_flatten_threshold_aft | (peaklet_classification_he,) | Threshold for AFT, above which we use a flatted boundary for rise timeSpecified values: (AFT bou... | <OMITTED> | (0.6, 100) |
| 12 | s1_max_rise_time_post100 | (peaklet_classification_he,) | Maximum S1 rise time for > 100 PE [ns] | <OMITTED> | 200 |
| 13 | s1_min_coincidence | (peaklet_classification_he,) | Minimum tight coincidence necessary to make an S1 | <OMITTED> | 2 |
Looks like we’re after the s1_min_coincidence option. Note this is not part of the event_basics data type, but of a data type called peak_classification. As you can see from the table, this option is not set in the current context, so the default value (3) is used.
To try out a different option, just pass it to get_df:
[30]:
# Let's use a short run for the following exaples
run_id = "038769"
[31]:
st2 = st.new_context()
st2.set_config(dict(s1_min_coincidence=50))
df_2 = st2.get_df(
run_id,
"event_info",
config=dict(s1_min_coincidence=50),
)
st2.event_scatter(run_id, events=df_2)
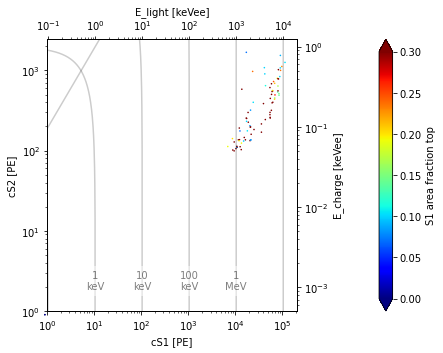
Notice all the small S1 events are indeed gone now.
Behind the scenes, this figured out which dataframes had to be remade: as it happens this time just event_basics and peak_basics. You will now have a new event_basics_<somehash> folder in ./custom_data which contains the results, as well as a new peak_basics_<somehash> folder.
More on configuration changes
Changing configuration can be done in two other ways. We can change it permanently in the current context:
st.set_config(dict(s1_min_coincidence=50))
Or we could make a new context, with this option set:
st_2 = st.new_context(config=dict(s1_min_coincidence=50))
(feeding it to get_df just does the latter behind the scenes).
If you just want to run a mini-analysis (like event_scatter), you can also pass a new config option directly to it, as in the example below.
Strax protects you from typos in the configuration. Suppose we typed s1_min_n_channelz instead:
[32]:
st.event_scatter(run_id, config=dict(s1_min_n_channelz=10), seconds_range=[0, 60])
Option s1_min_n_channelz not taken by any registered plugin
Option s1_min_n_channelz not taken by any registered plugin
---------------------------------------------------------------------------
DataNotAvailable Traceback (most recent call last)
Input In [32], in <module>
----> 1 st.event_scatter(run_id, config=dict(s1_min_n_channelz=10), seconds_range=[0,60])
File ~/software/dev_strax/straxen/straxen/mini_analysis.py:106, in mini_analysis.<locals>.decorator.<locals>.wrapped_f(context, run_id, **kwargs)
100 kwargs[dkind] = strax.apply_selection(
101 kwargs[dkind],
102 selection_str=kwargs['selection_str'],
103 time_range=kwargs['time_range'],
104 time_selection=kwargs['time_selection'])
105 else:
--> 106 kwargs[dkind] = context.get_array(
107 run_id,
108 dtypes,
109 selection_str=kwargs['selection_str'],
110 time_range=kwargs['time_range'],
111 time_selection=kwargs['time_selection'],
112 # Arguments for new context, if needed
113 config=kwargs.get('config'),
114 register=kwargs.get('register'),
115 storage=kwargs.get('storage', tuple()),
116 progress_bar=False,
117 )
119 # If user did not give time kwargs, but the function expects
120 # a time_range, try to add one based on the time range of the data
121 base_dkind = list(deps_by_kind.keys())[0]
File ~/software/dev_strax/strax/strax/context.py:1406, in Context.get_array(self, run_id, targets, save, max_workers, **kwargs)
1399 else:
1400 source = self.get_iter(
1401 run_ids[0],
1402 targets,
1403 save=save,
1404 max_workers=max_workers,
1405 **kwargs)
-> 1406 results = [x.data for x in source]
1408 results = np.concatenate(results)
1409 return results
File ~/software/dev_strax/strax/strax/context.py:1406, in <listcomp>(.0)
1399 else:
1400 source = self.get_iter(
1401 run_ids[0],
1402 targets,
1403 save=save,
1404 max_workers=max_workers,
1405 **kwargs)
-> 1406 results = [x.data for x in source]
1408 results = np.concatenate(results)
1409 return results
File ~/software/dev_strax/strax/strax/context.py:1234, in Context.get_iter(self, run_id, targets, save, max_workers, time_range, seconds_range, time_within, time_selection, selection_str, keep_columns, drop_columns, allow_multiple, progress_bar, _chunk_number, **kwargs)
1225 elif (self.context_config['timeout'] > 7200 or (
1226 self.context_config['allow_lazy'] and
1227 not self.context_config['allow_multiprocess'])):
1228 # For allow_multiple we don't want allow this when in lazy mode
1229 # with long timeouts (lazy-mode is disabled if multiprocessing
1230 # so if that is activated, we can also continue)
1231 raise RuntimeError(f'Cannot allow_multiple in lazy mode or '
1232 f'with long timeouts.')
-> 1234 components = self.get_components(run_id,
1235 targets=targets,
1236 save=save,
1237 time_range=time_range,
1238 chunk_number=_chunk_number)
1240 # Cleanup the temp plugins
1241 for k in list(self._plugin_class_registry.keys()):
File ~/software/dev_strax/strax/strax/context.py:979, in Context.get_components(self, run_id, targets, save, time_range, chunk_number)
975 savers = self._add_saver(savers, d_to_save, target_plugin,
976 _is_superrun, loading_this_data)
978 for target_i in targets:
--> 979 check_cache(target_i)
980 plugins = to_compute
982 intersec = list(plugins.keys() & loaders.keys())
File ~/software/dev_strax/strax/strax/context.py:894, in Context.get_components.<locals>.check_cache(target_i)
891 if target_plugin.save_when[target_i] == strax.SaveWhen.TARGET:
892 error_message += (f"\nFirst run st.make({run_id}, "
893 f"{target_i}) to make {target_i}.")
--> 894 raise strax.DataNotAvailable(error_message)
895 if '*' in self.context_config['forbid_creation_of']:
896 raise strax.DataNotAvailable(
897 f"{target_i} for {run_id} not found in any storage, and "
898 "your context specifies no new data can be created.")
DataNotAvailable: Time range selection assumes data is already available, but event_info for 038769 is not.
The result of get_df is just the same as if the option wasn’t set (just like in pax/hax), but you also get a warning about an unknown configuration option.
By the way, you can use
import warnings
warnings.filterwarnings("error")
to ensure any warning raises an exception instead.
Customization: new plugins
To add or change processing algorithms, or to define new variables to use in cuts, you have to write new strax plugins. These are somewhat similar to hax’s treemakers.
Suppose you have a brilliant new idea for peak classification. Strax does this in the peaklet_classification plugin, which produces:
[33]:
st.data_info("peaklet_classification")
[33]:
| Field name | Data type | Comment | |
|---|---|---|---|
| 0 | time | int64 | Start time since unix epoch [ns] |
| 1 | length | int32 | Length of the interval in samples |
| 2 | dt | int32 | Width of one sample [ns] |
| 3 | channel | int16 | Channel/PMT number |
| 4 | type | int8 | Classification of the peak(let) |
The first three fields contain time information of the peak. This is duplicated in many datatypes – unfortunately, this is necessary for strax to be able to track the data and combine it with other data. Instead of (time, length, dt), plugins could also provide (time, endtime). See here for more information.
The ‘channel’ field is an historical artifact.
The ‘type’ field contains the classification: 0 for unknown, 1 for S1, 2 for S2. (note #8)
You can find the original plugin in peaklet_processing.py Here’s how you would make a different classification plugin:
[34]:
import strax
import numpy as np
class AdvancedExpertClassification(strax.Plugin):
"""Everything is an S1!"""
# Name of the data type this plugin provides
provides = "peaklet_classification"
# Data types this plugin requires. Note we don't specify
# what plugins should produce them: maybe the default PeakBasics
# has been replaced by another AdvancedExpertBlabla plugin?
depends_on = ("peaklets",)
# Numpy datatype of the output
dtype = straxen.PeakletClassification.dtype
# Version of the plugin. Increment this if you change the algorithm.
__version__ = "0.0.2"
def compute(self, peaklets):
# Your code here.
# This function will be called several times with
# 'peaks' a numpy array of the datatype 'peaks'.
# Each time you'll see a small part of the run.
# You have to return a numpy array of the dtype you declared above
result = np.zeros(len(peaklets), self.dtype)
# Copy the basic time fields over from peaklets
for (_, field), _ in strax.time_dt_fields:
result[field] = peaklets[field]
# Store the classification results
# You might want to do real work here
result["type"] = 1
return result
# Instead of an array, you are also allowed to return a dictionary
# we can transform into an array.
# That is, (dict keys -> field names, values -> field values)
To use it in place of PeakClassification, we only have to register it. Again, we can do so permanently using
st.register(AdvancedExpertClassification)
or temporarily, by feeding the registration as an extra argument to get_df:
[35]:
df = st.get_df(
run_id,
"event_info",
register=AdvancedExpertClassification,
)
Found chunk without events?!
Found chunk without events?!
Found chunk without events?!
[36]:
df["s2_area"].max()
[36]:
nan
As you can see, all events are now S1-only events, as expected. Maybe this is not the best alternative classification :-)
This plugin was a rather basic plugin. You’ll also want to learn about LoopPlugins and OverlapWindowPlugins, but that’s beyond the scope of this tutorial.